2.3 Pandas
Contents
2.3 Pandas#
This section aims to provide new skills in python to handle structured, tabular data.
Learning outcome:
Manipulation of data frames (describing, filtering, …)
Learn about Lambda functions
Intro to datetime objects
Plotting data from data frames (histograms and maps)
Introduction to Plotly
Introduction to CSV & Parquet
This tutorial can be offered in a 2-hour course. Sections are labeled as Level 1, 2, 3 in Section 1, 2, 3, and instructors may choose to leave higher levels for asynchronous, self-guided learning.
We will work on several structured data sets: sensor metadata, seismic data product (earthquake catalog).
First, we import all the modules we need:
import numpy as np
import pandas as pd
import io
import requests
import matplotlib.pyplot as plt
from datetime import datetime, timedelta
This notebook uses plotly to make interactive plots
!pip install plotly
import plotly.express as px
import plotly.io as pio
# pio.renderers.default = 'vscode' # writes as standalone html,
# pio.renderers.default = 'iframe' # writes files as standalone html,
# pio.renderers.default = 'png' # writes files as standalone png,
# try notebook, jupyterlab, png, vscode, iframe
Requirement already satisfied: plotly in /Users/marinedenolle/opt/miniconda3/envs/mlgeo/lib/python3.9/site-packages (5.17.0)
Requirement already satisfied: tenacity>=6.2.0 in /Users/marinedenolle/opt/miniconda3/envs/mlgeo/lib/python3.9/site-packages (from plotly) (8.2.3)
Requirement already satisfied: packaging in /Users/marinedenolle/opt/miniconda3/envs/mlgeo/lib/python3.9/site-packages (from plotly) (23.2)
[notice] A new release of pip is available: 23.3.1 -> 24.2
[notice] To update, run: pip install --upgrade pip
1 Pandas Fundamentals#
1.1 Basics#
Pandas are composed of Series and DataFrame. Series are columns with attributes or keys. The DataFrame is a multi-dimensional table made up of Series.
We can create a DataFrame composed of series from scratch using Python dictionary:
data = {
'temperature' : [36,37,30,50],
'precipitation':[3,1,0,0]
}
my_pd = pd.DataFrame(data)
print(my_pd)
temperature precipitation
0 36 3
1 37 1
2 30 0
3 50 0
Each (key,value) item in the dataframe corresponds to a value in data. To get the keys of the dataframe, type:
my_pd.keys()
Index(['temperature', 'precipitation'], dtype='object')
get a specific Series (different from the array)
print(my_pd.temperature[:])
print(type(my_pd.temperature[:]))
0 36
1 37
2 30
3 50
Name: temperature, dtype: int64
<class 'pandas.core.series.Series'>
to get the value of a specific key (e.g., temperature), at a specific index (e.g., 2) type:
print(my_pd.temperature[2])
print(type(my_pd.temperature[2]))
30
<class 'numpy.int64'>
1.2 Reading a DataFrame from a CSV file#
We can read a pandas directly from a standard file. We will download a catalog of earthquakes
fname = ""
gist_dir = ""
!wget "https://raw.githubusercontent.com/UW-MLGEO/MLGeo-dataset/refs/heads/main/data/Global_Quakes_IRIS.csv"
# !wget "https://github.com/UW-MLGEO/MLGeo-dataset/blob/main/data/Global_Quakes_IRIS.csv"
ff = "Global_Quakes_IRIS.csv"
quake = pd.read_csv(ff)
--2024-10-07 05:13:23-- https://raw.githubusercontent.com/UW-MLGEO/MLGeo-dataset/refs/heads/main/data/Global_Quakes_IRIS.csv
Resolving raw.githubusercontent.com (raw.githubusercontent.com)... 185.199.111.133, 185.199.109.133, 185.199.110.133, ...
Connecting to raw.githubusercontent.com (raw.githubusercontent.com)|185.199.111.133|:443... connected.
HTTP request sent, awaiting response... 200 OK
Length: 129629 (127K) [text/plain]
Saving to: ‘Global_Quakes_IRIS.csv.1’
Global_Quakes_IRIS. 100%[===================>] 126.59K --.-KB/s in 0.05s
2024-10-07 05:13:23 (2.35 MB/s) - ‘Global_Quakes_IRIS.csv.1’ saved [129629/129629]
Now you use the head function to display what is in the file
# enter answer here
quake.head()
| time | latitude | longitude | depth | magnitude | description | |
|---|---|---|---|---|---|---|
| 0 | 2010-07-02 06:04:03.570 | -13.6098 | 166.6541 | 34400.0 | 6.3 | VANUATU ISLANDS |
| 1 | 2010-07-04 21:55:52.370 | 39.6611 | 142.5792 | 30100.0 | 6.3 | NEAR EAST COAST OF HONSHU |
| 2 | 2010-07-10 11:43:33.000 | 11.1664 | 146.0823 | 16900.0 | 6.3 | SOUTH OF MARIANA ISLANDS |
| 3 | 2010-07-12 00:11:20.060 | -22.2789 | -68.3159 | 109400.0 | 6.2 | NORTHERN CHILE |
| 4 | 2010-07-14 08:32:21.850 | -38.0635 | -73.4649 | 25700.0 | 6.6 | NEAR COAST OF CENTRAL CHILE |
Display the depth using two ways to use the pandas object
print(quake.depth)
print(quake['depth'])
0 34400.0
1 30100.0
2 16900.0
3 109400.0
4 25700.0
...
1780 10000.0
1781 105000.0
1782 608510.0
1783 10000.0
1784 35440.0
Name: depth, Length: 1785, dtype: float64
0 34400.0
1 30100.0
2 16900.0
3 109400.0
4 25700.0
...
1780 10000.0
1781 105000.0
1782 608510.0
1783 10000.0
1784 35440.0
Name: depth, Length: 1785, dtype: float64
Calculate basic statistic of the data using the function describe.
quake.describe()
| latitude | longitude | depth | magnitude | |
|---|---|---|---|---|
| count | 1785.000000 | 1785.000000 | 1785.000000 | 1785.000000 |
| mean | 0.840700 | 40.608674 | 82773.187675 | 6.382403 |
| std | 30.579308 | 125.558363 | 146988.302031 | 0.429012 |
| min | -69.782500 | -179.957200 | 0.000000 | 6.000000 |
| 25% | -19.905100 | -73.832200 | 12000.000000 | 6.100000 |
| 50% | -4.478900 | 113.077800 | 24400.000000 | 6.300000 |
| 75% | 27.161700 | 145.305700 | 58000.000000 | 6.600000 |
| max | 74.391800 | 179.998100 | 685500.000000 | 9.100000 |
Calculate mean and median of specific Series, for example depth.
# answer it here
print(quake.depth.mean())
print(quake.depth.median())
82773.18767507003
24400.0
1.3 Manipulating Pandas with Python#
Classic functions#
We will now practice how to modify the content of the DataFrame using functions. We will take the example where we want to change the depth values from meters to kilometers. First we can define this operation as a function
# this function converts a value in meters to a value in kilometers
m2km = 1000 # this is defined as a global variable
def meters2kilometers(x):
return x/m2km
# now test it using the first element of the quake DataFrame
meters2kilometers(quake.depth[0])
34.4
# this function converts a value in meters to a value in kilometers
def meters2kilometers2(x):
m2km = 1000 # this is defined as a global variable
return x/m2km
Let’s define another function that uses a local instead of global variable
# Apply the meters2kilometers2 function to the 'depth' column and add it as a new column 'depth_km' to the quake DataFrame
quake['depth_km'] = quake['depth'].apply(meters2kilometers2)
# Display the first few rows to verify the new column
quake.head()
| time | latitude | longitude | depth | magnitude | description | depth_km | |
|---|---|---|---|---|---|---|---|
| 0 | 2010-07-02 06:04:03.570 | -13.6098 | 166.6541 | 34400.0 | 6.3 | VANUATU ISLANDS | 34.4 |
| 1 | 2010-07-04 21:55:52.370 | 39.6611 | 142.5792 | 30100.0 | 6.3 | NEAR EAST COAST OF HONSHU | 30.1 |
| 2 | 2010-07-10 11:43:33.000 | 11.1664 | 146.0823 | 16900.0 | 6.3 | SOUTH OF MARIANA ISLANDS | 16.9 |
| 3 | 2010-07-12 00:11:20.060 | -22.2789 | -68.3159 | 109400.0 | 6.2 | NORTHERN CHILE | 109.4 |
| 4 | 2010-07-14 08:32:21.850 | -38.0635 | -73.4649 | 25700.0 | 6.6 | NEAR COAST OF CENTRAL CHILE | 25.7 |
Lambda functions#
We now discuss the lambda functions.
Lambda functions are small, anonymous functions in Python.
They are used for quick, simple operations without needing to define a full function using def.
In geoscience, where we deal with large datasets (e.g., climate data, seismic measurements), lambda functions allow us to process data efficiently.
Lambda functions in Pandas allow you to quickly transform, filter, or process this data with minimal code.
Please read additional tutorials from RealPython.
lambda x: x * 1.2
<function __main__.<lambda>(x)>
This lambda function multiplies any input x by 1.2, which could be useful for tasks like converting units (e.g., from km to m).
Below is an example on how to use lambda in a pandas dataframe.
Example 1:
The lambda function lambda x: (x - 32) * 5 / 9 converts Fahrenheit to Celsius for each value in the Temperature_F column.
import pandas as pd
# Sample DataFrame
data = {'Temperature_F': [32, 50, 77, 100]}
df = pd.DataFrame(data)
# Convert Fahrenheit to Celsius
df['Temperature_C'] = df['Temperature_F'].apply(lambda x: (x - 32) * 5 / 9)
print(df)
Temperature_F Temperature_C
0 32 0.000000
1 50 10.000000
2 77 25.000000
3 100 37.777778
# Now the equivalent in lambda is:
lambda_meters2kilometers = lambda x:x/1000
# x is the variable
Now apply the function to the entire series
Student response section
This section is left for the student to complete.
# apply it to the entire series
lambda_meters2kilometers(quake.depth)
0 34.40
1 30.10
2 16.90
3 109.40
4 25.70
...
1780 10.00
1781 105.00
1782 608.51
1783 10.00
1784 35.44
Name: depth, Length: 1785, dtype: float64
Lambda functions can take several inputs
# you can add several variables into lambda functions
remove_anything = lambda x,y:x-y
remove_anything(3,2)
1
This did not affect the values of the DataFrame, check it:
quake.depth
0 34400.0
1 30100.0
2 16900.0
3 109400.0
4 25700.0
...
1780 10000.0
1781 105000.0
1782 608510.0
1783 10000.0
1784 35440.0
Name: depth, Length: 1785, dtype: float64
Instead, you could overwrite quake.depth=X. Try two approaches but just do it once! You can use python functions map (a function to apply functions that is generic to Python) and apply (a function to apply functions that is specific to Pandas).
Student response section
This section is left for the student to complete.
Try map to apply a lambda function that rescale depth from meters to kilometers.
#type answer below
quake.depth=quake.depth.map(lambda x:x/1000)
Discuss in class: What happened to the depth field?
What happened to the original data frame?
Try apply to apply a lambda function that rescale depth from meters to kilometers.
# or like this
quake.depth=quake.depth.apply(lambda x:x/1000)
Plot a histogram of the depth distributions using matplotlib function hist.
Student response section
This section is left for the student to complete.
# answer here
plt.hist(quake.depth,100)
plt.grid(True)
plt.xlabel('Quake depth (km)')
plt.show()
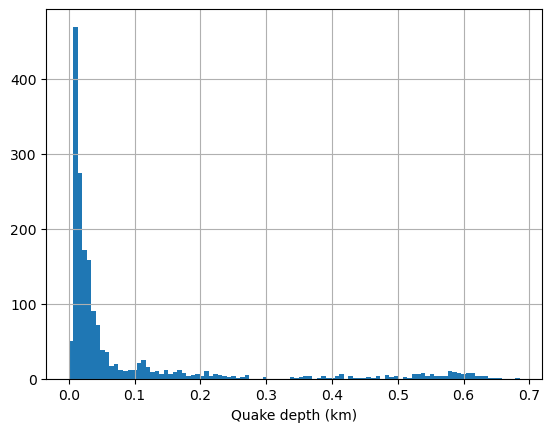
You can use the interactive plotting package Plotly. First we will show a histogram of the event depth using the function histogram.
fig = px.histogram(quake, #specify what dataframe to use
x="depth", #specify the variable for the histogram
nbins=50, #number of bins for the histogram
height=400, #dimensions of the figure
width=600);
fig.show()
Example 2: Conditional Logic for Earthquake Magnitude#
You can use lambda functions to add conditional logic, such as classifying earthquake magnitudes into categories.
Scenario: You have a column Magnitude containing seismic event magnitudes, and you want to classify events as “Minor”, “Moderate”, or “Severe”.
# Classify earthquake magnitudes
quake['Category'] = quake['magnitude'].apply(lambda x: 'Minor' if x < 4.0 else ('Moderate' if x < 6.5 else 'Severe'))
quake.head()
| time | latitude | longitude | depth | magnitude | description | depth_km | Category | |
|---|---|---|---|---|---|---|---|---|
| 0 | 2010-07-02 06:04:03.570 | -13.6098 | 166.6541 | 0.0344 | 6.3 | VANUATU ISLANDS | 34.4 | Moderate |
| 1 | 2010-07-04 21:55:52.370 | 39.6611 | 142.5792 | 0.0301 | 6.3 | NEAR EAST COAST OF HONSHU | 30.1 | Moderate |
| 2 | 2010-07-10 11:43:33.000 | 11.1664 | 146.0823 | 0.0169 | 6.3 | SOUTH OF MARIANA ISLANDS | 16.9 | Moderate |
| 3 | 2010-07-12 00:11:20.060 | -22.2789 | -68.3159 | 0.1094 | 6.2 | NORTHERN CHILE | 109.4 | Moderate |
| 4 | 2010-07-14 08:32:21.850 | -38.0635 | -73.4649 | 0.0257 | 6.6 | NEAR COAST OF CENTRAL CHILE | 25.7 | Severe |
We will now make a new plot of the location of the earthquakes. We will use Plotly tool.
The markersize will be scaled with the earthquake magnitude. To do so, we add a marker_size series in the DataFrame
quake['marker_size'] = quake['magnitude'].apply(lambda x: np.fix(np.exp(x))) # add marker size as exp(mag)
quake['magnitude_bin'] = quake['magnitude'].apply(lambda x: 0.5*np.fix(2*x)) # add marker size as exp(mag)
# another way to do it
quake['marker_size'] = np.fix(np.exp(quake['magnitude'])) # add marker size as exp(mag)
quake['magnitude_bin'] = 0.5*np.fix(2*quake['magnitude']) # add marker size as exp(mag)
1.4 Intermediate Manipulation with Pandas#
You can also apply lambda functions to work with multiple columns, which is common in geoscientific datasets, where you might have spatial or temporal data.
Example 3: Calculating an Index Based on Multiple Measurements
Scenario: You have rainfall (Rainfall_mm) and evaporation (Evaporation_mm) data, and you want to calculate the Net Water Balance for each record.
data = {'Rainfall_mm': [100, 80, 120], 'Evaporation_mm': [60, 70, 65]}
df = pd.DataFrame(data)
# Calculate Net Water Balance
df['Net_Water_Balance'] = df.apply(lambda row: row['Rainfall_mm'] - row['Evaporation_mm'], axis=1)
print(df)
Rainfall_mm Evaporation_mm Net_Water_Balance
0 100 60 40
1 80 70 10
2 120 65 55
In this example, lambda row: row['Rainfall_mm'] - row['Evaporation_mm'] calculates the net water balance by subtracting evaporation from rainfall for each record.
1.5 Advanced: Time Series Data Manipulation with Lambda Functions#
Geoscientists frequently work with time series data (e.g., climate data). Lambda functions can be used for efficient data transformations within time series.
Example 4: Applying a Rolling Window Calculation Scenario: Suppose you have daily temperature data, and you want to calculate a 7-day rolling average.
# Sample daily temperature data
data = {'Date': pd.date_range(start='2023-09-01', periods=10, freq='D'),
'Temperature_C': [20, 22, 23, 21, 19, 24, 25, 26, 22, 20]}
df = pd.DataFrame(data)
# Calculate 7-day rolling average using lambda
df['7_day_avg'] = df['Temperature_C'].rolling(window=3).apply(lambda x: x.mean())
df.head()
| Date | Temperature_C | 7_day_avg | |
|---|---|---|---|
| 0 | 2023-09-01 | 20 | NaN |
| 1 | 2023-09-02 | 22 | NaN |
| 2 | 2023-09-03 | 23 | 21.666667 |
| 3 | 2023-09-04 | 21 | 22.000000 |
| 4 | 2023-09-05 | 19 | 21.000000 |
Here, lambda x: x.mean() calculates the rolling mean over a 3-day window for temperature data, which is essential in climate analysis for smoothing short-term fluctuations.
1.6 Aggregate function#
The agg is a powerful method in Pandas that allows you to perform multiple operations on DataFrames and Series. You can apply one or more aggregation functions such as sum, mean, min, max, etc., on different columns or groups of data.
# Sample DataFrame
data = {
'A': [1, 2, 3, 4],
'B': [5, 6, 7, 8]
}
df = pd.DataFrame(data)
# Apply aggregation functions
result = df['A'].agg(['sum', 'mean'])
print(result)
sum 10.0
mean 2.5
Name: A, dtype: float64
Aggregating Multiple Columns with Multiple Functions#
You can pass a dictionary to agg where the keys are column names and the values are the functions to be applied.
# Apply different functions to different columns
result = df.agg({'A': ['sum', 'mean'], 'B': ['min', 'max']})
print(result)
A B
sum 10.0 NaN
mean 2.5 NaN
min NaN 5.0
max NaN 8.0
In this example, column ‘A’ gets sum and mean, while column ‘B’ gets min and max.
You may also use costum functions
# Custom function to calculate range (max - min)
def data_range(x):
return x.max() - x.min()
# Apply custom function
result = df.agg({'A': ['mean', data_range], 'B': data_range})
print(result)
A B
mean 2.5 NaN
data_range 3.0 3.0
2.4 Mapping using Plotly#
Now we will plot the earthquakes locations on a map using the Plotly package. More tutorials on Plotly. The input of the function is self-explanatory and typical of Python’s function. The code documentation of Plotly scatter_geo lists the variables.
fig = px.scatter_geo(quake,
lat='latitude',lon='longitude',
range_color=(6,9),
height=600, width=600,
size='marker_size', color='magnitude',
hover_name="description",
hover_data=['description','magnitude','depth']);
fig.update_geos(resolution=110, showcountries=True)
fig.update_geos(resolution=110, showcountries=True,projection_type="orthographic")
fig
The lambda function lambda x: 'Minor' if x < 4.0 else ('Moderate' if x < 6.5 else 'Severe') classifies earthquake magnitudes based on their values.
The data was sorted by time. We now want to sort and show the data instead by magnitude. We use the pandas function sort to create a new DataFrame with sorted values.
quakes2plot=quake.sort_values(by='magnitude_bin')
quakes2plot.head()
| time | latitude | longitude | depth | magnitude | description | depth_km | Category | marker_size | magnitude_bin | |
|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 2010-07-02 06:04:03.570 | -13.6098 | 166.6541 | 0.03440 | 6.3 | VANUATU ISLANDS | 34.40 | Moderate | 544.0 | 6.0 |
| 1112 | 2017-08-31 17:06:55.750 | -1.1590 | 99.6881 | 0.04314 | 6.3 | SOUTHERN SUMATRA | 43.14 | Moderate | 544.0 | 6.0 |
| 1111 | 2017-08-27 04:17:51.010 | -1.4513 | 148.0803 | 0.00800 | 6.3 | ADMIRALTY ISLANDS REGION | 8.00 | Moderate | 544.0 | 6.0 |
| 1110 | 2017-08-19 02:00:52.550 | -17.9609 | -178.8406 | 0.54400 | 6.4 | FIJI ISLANDS REGION | 544.00 | Moderate | 601.0 | 6.0 |
| 1108 | 2017-08-13 03:08:10.560 | -3.7682 | 101.6228 | 0.03100 | 6.4 | SOUTHERN SUMATRA | 31.00 | Moderate | 601.0 | 6.0 |
Now we will plot again using Plotly
fig = px.scatter_geo(quakes2plot,
lat='latitude',lon='longitude',
range_color=(6,9),
height=600, width=600,
size='marker_size', color='magnitude',
hover_name="description",
hover_data=['description','magnitude','depth']);
fig.update_geos(resolution=110, showcountries=True)
# fig.update_geos(resolution=110, showcountries=True,projection_type="orthographic")
3 Create a Pandas from a generic text file.#
The python package pandas is very useful to read csv files, but also many text files that are more or less formatted as one observation per row and one column for each feature.
As an example, we are going to look at the list of seismic stations from the Northern California seismic network, available here:
url = 'http://ncedc.org/ftp/pub/doc/NC.info/NC.channel.summary.day'
# this gets the file linked in the URL page and convert it to a string
s = requests.get(url).content
# this will convert the string, decode it , and make it a table
data = pd.read_csv(io.StringIO(s.decode('utf-8')), header=None, skiprows=2, sep='\s+', usecols=list(range(0, 13)))
# because columns/keys were not assigned, assign them now
data.columns = ['station', 'network', 'channel', 'location', 'rate', 'start_time', 'end_time', 'latitude', 'longitude', 'elevation', 'depth', 'dip', 'azimuth']
Let us look at the data. They are now stored into a pandas dataframe.
data.head()
| station | network | channel | location | rate | start_time | end_time | latitude | longitude | elevation | depth | dip | azimuth | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | AAR | NC | EHZ | -- | 0.0 | 1976/07/20,17:38:00 | 1977/12/01,22:37:00 | 39.27594 | -121.02696 | 911.0 | 0.0 | -90.0 | 0.0 |
| 1 | AAR | NC | EHZ | -- | 0.0 | 1977/12/01,22:37:00 | 1981/06/12,19:15:00 | 39.27594 | -121.02696 | 911.0 | 0.0 | -90.0 | 0.0 |
| 2 | AAR | NC | EHZ | -- | 0.0 | 1981/06/12,19:15:00 | 1984/01/01,00:00:00 | 39.27594 | -121.02696 | 911.0 | 0.0 | -90.0 | 0.0 |
| 3 | AAR | NC | EHZ | -- | 100.0 | 1984/01/01,00:00:00 | 1985/11/03,22:00:00 | 39.27594 | -121.02696 | 911.0 | 0.0 | -90.0 | 0.0 |
| 4 | AAR | NC | EHZ | -- | 100.0 | 1985/11/03,22:00:00 | 1987/03/10,23:15:00 | 39.27594 | -121.02696 | 911.0 | 0.0 | -90.0 | 0.0 |
We can output the first element of the DataFrame:
data.iloc[0]
station AAR
network NC
channel EHZ
location --
rate 0.0
start_time 1976/07/20,17:38:00
end_time 1977/12/01,22:37:00
latitude 39.27594
longitude -121.02696
elevation 911.0
depth 0.0
dip -90.0
azimuth 0.0
Name: 0, dtype: object
# display the type of each column
data.dtypes
station object
network object
channel object
location object
rate float64
start_time object
end_time object
latitude float64
longitude float64
elevation float64
depth float64
dip float64
azimuth float64
dtype: object
data.iloc[:, 0]
0 AAR
1 AAR
2 AAR
3 AAR
4 AAR
...
29434 WMP
29435 WMP
29436 WMP
29437 WSL
29438 WWVB
Name: station, Length: 29439, dtype: object
The start_time and end_time are stored as object or string, convert these columns to datetime objects. With a too-simple prompt, CoPilot may fail at converting:
# convert start_time and end_time to datetime
data['start_time'] = pd.to_datetime(data['start_time'])
data['end_time'] = pd.to_datetime(data['end_time'])
/var/folders/js/lzmy975n0l5bjbmr9db291m00000gn/T/ipykernel_26544/3238846712.py:2: UserWarning:
Could not infer format, so each element will be parsed individually, falling back to `dateutil`. To ensure parsing is consistent and as-expected, please specify a format.
---------------------------------------------------------------------------
DateParseError Traceback (most recent call last)
Cell In[78], line 2
1 # convert start_time and end_time to datetime
----> 2 data['start_time'] = pd.to_datetime(data['start_time'])
3 data['end_time'] = pd.to_datetime(data['end_time'])
File ~/opt/miniconda3/envs/mlgeo/lib/python3.9/site-packages/pandas/core/tools/datetimes.py:1108, in to_datetime(arg, errors, dayfirst, yearfirst, utc, format, exact, unit, infer_datetime_format, origin, cache)
1106 result = arg.tz_localize("utc")
1107 elif isinstance(arg, ABCSeries):
-> 1108 cache_array = _maybe_cache(arg, format, cache, convert_listlike)
1109 if not cache_array.empty:
1110 result = arg.map(cache_array)
File ~/opt/miniconda3/envs/mlgeo/lib/python3.9/site-packages/pandas/core/tools/datetimes.py:254, in _maybe_cache(arg, format, cache, convert_listlike)
252 unique_dates = unique(arg)
253 if len(unique_dates) < len(arg):
--> 254 cache_dates = convert_listlike(unique_dates, format)
255 # GH#45319
256 try:
File ~/opt/miniconda3/envs/mlgeo/lib/python3.9/site-packages/pandas/core/tools/datetimes.py:490, in _convert_listlike_datetimes(arg, format, name, utc, unit, errors, dayfirst, yearfirst, exact)
487 if format is not None and format != "mixed":
488 return _array_strptime_with_fallback(arg, name, utc, format, exact, errors)
--> 490 result, tz_parsed = objects_to_datetime64ns(
491 arg,
492 dayfirst=dayfirst,
493 yearfirst=yearfirst,
494 utc=utc,
495 errors=errors,
496 allow_object=True,
497 )
499 if tz_parsed is not None:
500 # We can take a shortcut since the datetime64 numpy array
501 # is in UTC
502 dta = DatetimeArray(result, dtype=tz_to_dtype(tz_parsed))
File ~/opt/miniconda3/envs/mlgeo/lib/python3.9/site-packages/pandas/core/arrays/datetimes.py:2346, in objects_to_datetime64ns(data, dayfirst, yearfirst, utc, errors, allow_object)
2343 # if str-dtype, convert
2344 data = np.array(data, copy=False, dtype=np.object_)
-> 2346 result, tz_parsed = tslib.array_to_datetime(
2347 data,
2348 errors=errors,
2349 utc=utc,
2350 dayfirst=dayfirst,
2351 yearfirst=yearfirst,
2352 )
2354 if tz_parsed is not None:
2355 # We can take a shortcut since the datetime64 numpy array
2356 # is in UTC
2357 # Return i8 values to denote unix timestamps
2358 return result.view("i8"), tz_parsed
File tslib.pyx:403, in pandas._libs.tslib.array_to_datetime()
File tslib.pyx:552, in pandas._libs.tslib.array_to_datetime()
File tslib.pyx:517, in pandas._libs.tslib.array_to_datetime()
File conversion.pyx:546, in pandas._libs.tslibs.conversion.convert_str_to_tsobject()
File parsing.pyx:331, in pandas._libs.tslibs.parsing.parse_datetime_string()
File parsing.pyx:660, in pandas._libs.tslibs.parsing.dateutil_parse()
DateParseError: Unknown datetime string format, unable to parse: 1976/07/20,17:38:00, at position 0
data['startdate'] = pd.to_datetime(data['start_time'], format='%Y/%m/%d,%H:%M:%S')
data.head()
| station | network | channel | location | rate | start_time | end_time | latitude | longitude | elevation | depth | dip | azimuth | startdate | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | AAR | NC | EHZ | -- | 0.0 | 1976/07/20,17:38:00 | 1977/12/01,22:37:00 | 39.27594 | -121.02696 | 911.0 | 0.0 | -90.0 | 0.0 | 1976-07-20 17:38:00 |
| 1 | AAR | NC | EHZ | -- | 0.0 | 1977/12/01,22:37:00 | 1981/06/12,19:15:00 | 39.27594 | -121.02696 | 911.0 | 0.0 | -90.0 | 0.0 | 1977-12-01 22:37:00 |
| 2 | AAR | NC | EHZ | -- | 0.0 | 1981/06/12,19:15:00 | 1984/01/01,00:00:00 | 39.27594 | -121.02696 | 911.0 | 0.0 | -90.0 | 0.0 | 1981-06-12 19:15:00 |
| 3 | AAR | NC | EHZ | -- | 100.0 | 1984/01/01,00:00:00 | 1985/11/03,22:00:00 | 39.27594 | -121.02696 | 911.0 | 0.0 | -90.0 | 0.0 | 1984-01-01 00:00:00 |
| 4 | AAR | NC | EHZ | -- | 100.0 | 1985/11/03,22:00:00 | 1987/03/10,23:15:00 | 39.27594 | -121.02696 | 911.0 | 0.0 | -90.0 | 0.0 | 1985-11-03 22:00:00 |
# do the same for end times
# Avoid 'OutOfBoundsDatetime' error with year 3000
enddate = data['end_time'].str.replace('3000', '2026')
enddate = pd.to_datetime(enddate, format='%Y/%m/%d,%H:%M:%S')
data['enddate'] = enddate
data.head()
| station | network | channel | location | rate | start_time | end_time | latitude | longitude | elevation | depth | dip | azimuth | startdate | enddate | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | AAR | NC | EHZ | -- | 0.0 | 1976/07/20,17:38:00 | 1977/12/01,22:37:00 | 39.27594 | -121.02696 | 911.0 | 0.0 | -90.0 | 0.0 | 1976-07-20 17:38:00 | 1977-12-01 22:37:00 |
| 1 | AAR | NC | EHZ | -- | 0.0 | 1977/12/01,22:37:00 | 1981/06/12,19:15:00 | 39.27594 | -121.02696 | 911.0 | 0.0 | -90.0 | 0.0 | 1977-12-01 22:37:00 | 1981-06-12 19:15:00 |
| 2 | AAR | NC | EHZ | -- | 0.0 | 1981/06/12,19:15:00 | 1984/01/01,00:00:00 | 39.27594 | -121.02696 | 911.0 | 0.0 | -90.0 | 0.0 | 1981-06-12 19:15:00 | 1984-01-01 00:00:00 |
| 3 | AAR | NC | EHZ | -- | 100.0 | 1984/01/01,00:00:00 | 1985/11/03,22:00:00 | 39.27594 | -121.02696 | 911.0 | 0.0 | -90.0 | 0.0 | 1984-01-01 00:00:00 | 1985-11-03 22:00:00 |
| 4 | AAR | NC | EHZ | -- | 100.0 | 1985/11/03,22:00:00 | 1987/03/10,23:15:00 | 39.27594 | -121.02696 | 911.0 | 0.0 | -90.0 | 0.0 | 1985-11-03 22:00:00 | 1987-03-10 23:15:00 |
Use Plotly to map the stations.
data.dropna(inplace=True)
data=data[data.longitude!=0]
fig = px.scatter_geo(data,
lat='latitude',lon='longitude',
range_color=(6,9),
height=600, width=600,
hover_name="station",
hover_data=['network','station','channel','rate']);
fig.update_geos(resolution=110, showcountries=True)
fig = px.scatter_mapbox(data,
lat='latitude',lon='longitude',
range_color=(6,9),mapbox_style="carto-positron",
height=600, width=500,
hover_name="station",
hover_data=['network','station','channel','rate']);
fig.update_layout(title="Northern California Seismic Network")
fig.show()
4: Exercise#
We will now practice on manipulating pandas. This exercise pulls station metadata from a URL and students are expected to practice on specific tasks.
Download data from the NCEDC URL
url = 'http://ncedc.org/ftp/pub/doc/NC.info/NC.channel.summary.day'
The column names are shown at the top of the text file, make a list of strings of these names in order to rename the columns once the dataframe is made
# students answer here
# students answer here
# request the data from the URL and use the IO package
# assign the column names
data.columns = ['station', 'network', 'channel', 'location', 'rate', 'start_time', 'end_time', 'latitude', 'longitude', 'elevation', 'depth', 'dip', 'azimuth']
Find the row of channel KCPB
# find the row of station KCPB
Now select two stations of your choice using |.
# answer below
Now select a given station and a specific channel code, example is KCPB and channel code HNZ.
# Select two stations, use the typical "AND"
You may also choose the pandas function isin to select rows that have a given attribute that belongs to a list. For instance, use isin to select all rows that have the key station within a list ['KCBP','KHBB' ].
# students answer here
Q Use panda native functions to calculate how many unique sites (station names) there are in the network.
# students answer here
Q Use pandas native functions to select the unique set of channel codes that end with Z: this will tell you of how many types digitized data the seismic network manages.
# students answer here
Q What station names has the most number of channels? hint you may use value_counts().
# students answer here
Q What is the maximum difference in elevation between the stations using lambda functions
# students answer here
Here, pandas does not recognize the start_time and end_time columns as a datetime format, so we cannot use datetime operations on them. We first need to convert these columns into a datetime format:
# answer here
# Transform column from string into datetime format
# do the same for end times
We can now look when each seismic station was installed using groupby and sorting by the earliest deployment (i.e., the minimum of the start_time)
# students answer here
Select the stations that were deployed first and recovered last using agg and lambda functions
# answer here
4. CSV vs Parquet#
Parquet is a compressed data format that stores and compresses the columns. It is fast for I/O and compact format.
Save data into a CSV file:
%timeit data.to_csv("my_metadata.csv")
!ls -lh my_metadata.csv
195 ms ± 1.29 ms per loop (mean ± std. dev. of 7 runs, 10 loops each)
-rw-r--r--@ 1 marinedenolle staff 4.2M Oct 7 05:15 my_metadata.csv
Try and save in Parquet and compare time and memory.
%timeit data.to_parquet("my_metadata.pq")
!ls -lh my_metadata.pq
25.2 ms ± 10.8 ms per loop (mean ± std. dev. of 7 runs, 1 loop each)
-rw-r--r--@ 1 marinedenolle staff 591K Oct 7 05:15 my_metadata.pq
